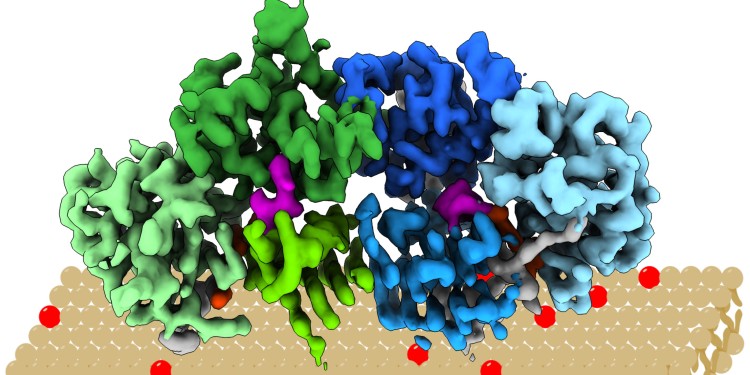
First 3D structure of regulator protein revealed
Proteins are indispensable components in living organisms. They are not only “building material” for the body – they also make molecular communication between cells possible, they are needed for nerve impulses to occur, and they control chemical reactions. What is decisive for proteins to function is their three-dimensional structure. If this is known, conclusions can be drawn about how proteins function. A team of researchers led by Prof. Daniel Kümmel from the University of Münster and Prof. Stefan Raunser from the Max Planck Institute (MPI) of Molecular Physiology in Dortmund has now clarified the structure of a protein complex which is an important regulator of cellular degradation processes.
The protein complex “Mon1/Ccz1” determines which intracellular vesicles deliver their content to the cellular “recycling centre”, the lysosome. To this end, it docks onto the vesicle membrane, where it introduces a label. Intracellular vesicles are membrane bubbles which transport material through the cell. In the lysosome, the content is degraded and re-used. By elucidating the structure in almost atomic resolution, the researchers were now able to clarify, among other things, how the protein complex recognises the appropriate vesicles. For example, they showed that the complex has a positively charged and relatively flat area which determines its orientation after docking onto the vesicle membrane.
“Mon1/Ccz1” belongs to a family of regulators for which no structural information exists. These complexes are involved in a range of cellular processes and are sometimes associated with the occurrence of developmental disorders such as albinism and blood clotting disorders. “Our structure now provides a basis for a better understanding of these connections,” says Daniel Kümmel.
The protein complex examined comes from the filamentous fungus Chaetomium thermophilum and is particularly stable and easy to handle under laboratory conditions. It can serve as a model for human proteins. In order to determine the protein’s structure, the researchers used high-resolution cryogenic electron microscopy. "With this method, we can study the structure of proteins at temperatures around minus 150 degrees Celsius in an almost natural state," says Stefan Raunser.
The researchers checked their results by means of biochemical studies, for example sedimentation assays. In this case, the protein-membrane interaction is demonstrated with artificial vesicles and purified protein in vitro, i.e. outside the organism. "The structure of Mon1/Ccz1 has a unique architecture that, to our knowledge, has not been demonstrated in any other protein complex. It could serve as a blueprint for a better understanding of other related regulatory proteins. We want to continue our successful collaboration," Daniel Kümmel and Stefan Raunser agree.
Further information and funding
The study was published in the interdisciplinary journal "Proceedings of the National Academy of Sciences of the United States of America". In addition to scientists from WWU Münster and MPI Dortmund, researchers from the University of Osnabrück were also involved. The German Research Foundation provided financial support for the work as part of Collaborative Research Centre (CRC) 944.
Original publication
Björn U. Klink, Eric Herrmann, Claudia Antonia, Lars Langemeyer, Stephan Kiontke, Christos Gatsogiannis, Christian Ungermann, Stefan Raunser, Daniel Kümmel: Structure of the Mon1-Ccz1 complex reveals molecular basis of membrane binding for Rab7 activation. PNAS February 8, 2022 119 (6), DOI: 10.1073/pnas.2121494119
