










The Institute of Integrative Cell Biology and Physiology (IIZP) was formed in 2021 by combining the former Institute of Molecular Cell Biology and the Institute of Zoophysiology.
Our institute embodies the close connection between physiological and cell biological processes that cannot be fully understood in isolation. At the IIZP, our mission is to explore the complex structures and physiological processes in animal cells and whole organisms across a range of time and length scales. Following this approach, our aim is to develop a comprehensive understanding of the molecular, cellular and biophysical principles of developmentally, physiologically and pathologically relevant processes.
The research activities at the IIZP integrate experiments on individual molecules, isolated cells, tissues and living organisms. We are using a wide range of model systems that include cell cultures, amoebae, crustaceans, nematodes, fruit flies as well as mouse models. For our integrative approach, we combine biochemical and genetic technologies with high-resolution, quantitative microscopy approaches.
Please visit our research groups for more information.
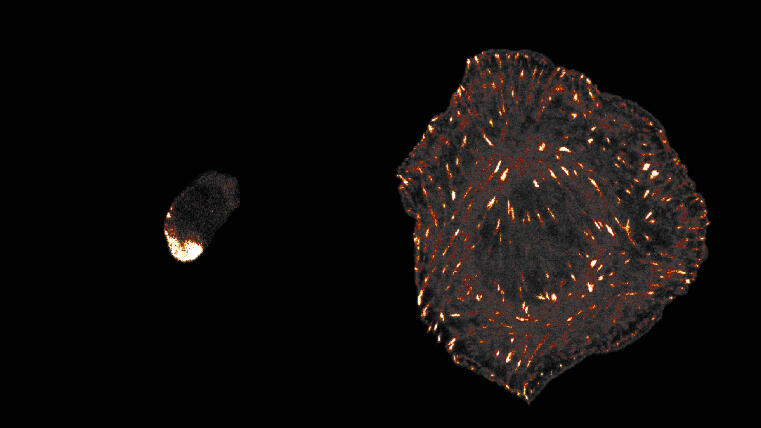
Biologists at the University of Münster discover evolutionary origin of animal cell adhesion / ‘Talin’ protein plays a central role
Cells of all animals – including humans – are characterized by their ability to adhere particularly well to surfaces in their environment. This mechanically stable adhesion enables the development of complex tissues and organs and is made possible by cell surface receptors called integrins. However, it is unclear how this form of cell adhesion developed over the course of evolution, as many single-celled organisms do not have integrin receptors. A team led by Prof. Carsten Grashoff and doctoral student Srishti Rangarajan from the Institute of Integrative Cell Biology and Physiology at the University of Münster has now shown that the talin protein plays a central and evolutionarily conserved role in cell adhesion.
Background and method
Talin is found in numerous eukaryotic (nucleus containing) single-celled organisms and also in all animal cells. There, the protein ensures the mechanical attachment of integrins to the cell interior. Through comparative studies in amoebae and animal cells, the researchers have now demonstrated that talin – much like in humans – also transmits mechanical forces during cell adhesion in single-celled organisms.
Even though each molecule only bears a force of a few trillionths of a Newton, this mechanical function of talin appears to be crucial for successful cell adhesion. The talin protein has a number of additional tasks in human cells that have not yet been observed in amoebae. However, the crucial mechanical role of the protein had probably already developed long before the first animals appeared.
Srishti Rangarajan emphasises: “The integrin-mediated adhesion of animal cells is described in all modern textbooks on cell biology. However, it appears to be merely a specialisation of a much older cell adhesion mechanism that originated in single-celled organisms and is mediated by talin.”
The team used methods from molecular genetics, high-resolution fluorescence microscopy and molecular force microscopy measurements.
Funding
The Volkswagen Foundation provided financial support for the work.
Original publication
Srishti Rangarajan, Lena Espeter, Hannes C.A. Drexler, Anna Chrostek-Grashoff and Carsten Grashoff. Talin force coupling underlies eukaryotic cell-substrate adhesion. Nat Commun. 2025 Dec 6;16(1):10950. doi: 10.1038/s41467-025-67354-8
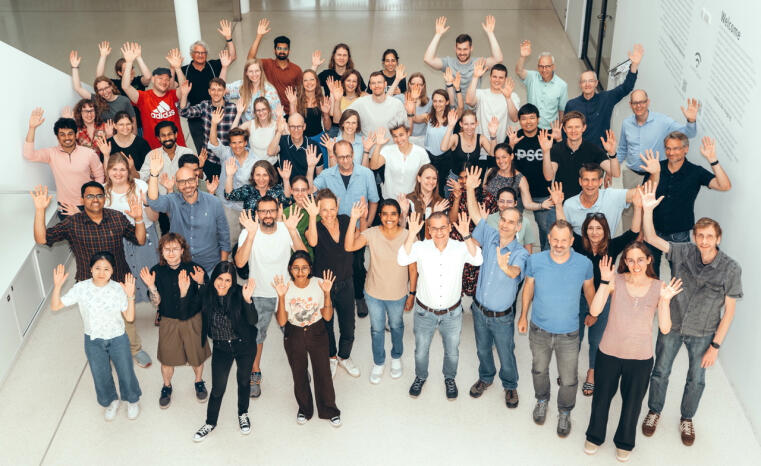


On November 21, the German Research Foundation (DFG) announced that the Collaborative Research Center (SFB) “Dynamic Cellular Interfaces: Formation and Function” will receive a further four years of funding totaling 12 million Euros.
The research network, coordinated by Prof. Stefan Luschnig from the Institute of Integrative Cell Biology and Physiology (IIZP), can thus continue its successful work on fundamental questions of cell and developmental biology as well as biomedicine. The research team investigates how cells communicate to exchange information, substances, and mechanical signals at the cell surface. These dynamic cellular interfaces control signal processing and development and are therefore of central importance for the orderly architecture of tissues and the normal function of organs. To understand these fundamental processes, the researchers are combining their expertise from various disciplines, including computer modeling, organic chemistry, structural biology, genetics, and molecular cell biology. Twenty-seven scientists from the Faculties of Biology, Chemistry, and Medicine at the University of Münster are involved in the consortium. They are working closely with the Max Planck Institute for Molecular Biomedicine and a sub-project at the Technical University of Dortmund.

The IIZP welcomes Dr. Maik Bischoff, who has joined the institute in October 2025 to establish his independent junior research group. His team investigates how organs acquire their shape and chirality, with a particular interest in the role of mesenchymal cell layers, such as smooth musculature, in sculpting organ architecture.
Using Drosophila melanogaster as a genetically accessible model, the group studies how collective cell migration, mechanosensation, and contact-based interactions drive complex morphogenetic processes. Their work combines high-resolution live-cell imaging, quantitative image analysis, genetics, and molecular biology to uncover how dynamic behaviors of individual cells translate into emergent, tissue-scale patterns.
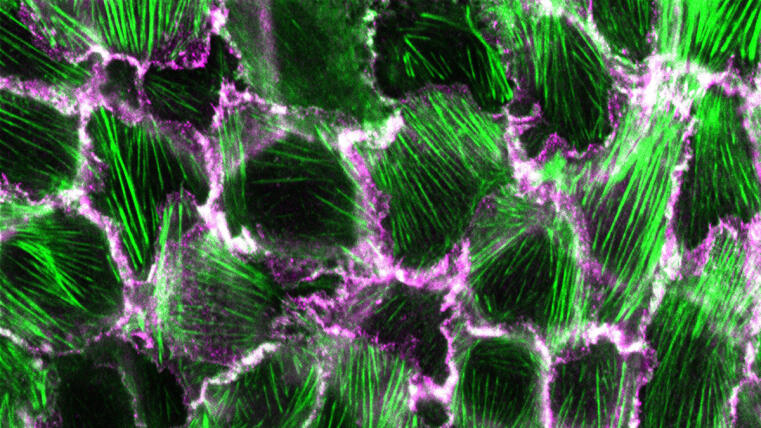
Researchers at the University of Münster develop method to visualize structural changes in a prominent adhesion protein / Study published in Communications Biology
The formation and maintenance of epithelia is crucial for the development and survival of all animals. Cadherin-based complexes, known as adhesion junctions, are essential for the integrity of these tissues, forming robust yet dynamic cell-cell adhesions. However, the molecular details underlying the formation of these important structures are not fully understood. A new study by the Grashoff group at the IIZP sheds light on this fundamental cell biological process.
Background and method
In their open-access study recently published in Communications Biology, the authors show that the maturation of adhesion junctions—found, for example, in human skin and intestines—is accompanied by a structural change within a protein that has previously been shown to be essential for the formation of cell-cell contacts in animals: α-catenin. By combining a novel α-catenin biosensor with fluorescence lifetime and anisotropy imaging, the authors show that the molecule undergoes a critical conformational change in its C-terminal actin-binding domain when adhesion junctions mature. Surprisingly, this conformational change, which is believed to strengthen cell-cell connections, correlates with increased protein turnover. The identified mechanism may therefore explain why epithelial tissues can form mechanically stable yet dynamic structures.
Funding
This work was funded by the German Research Foundation (DFG).
Original publication
Lukas Windgasse and Carsten Grashoff. A conformational change in α-catenin’s actin-binding domain governs adherens junction maturation. Commun Biol 8, 1325 (2025). DOI: 10.1038/s42003-025-08785-3