Cellular membrane and lipid dynamics
Cellular membranes represent highly dynamic structures that are crucially involved in almost all cellular responses to environmental cues. Membranes are a composite of lipids and proteins that are engaged in multiple dynamic interactions and can form transient domains of nano- to microscale dimensions. These membrane microdomains are thought to actively participate in many membrane-related events including signaling across the plasma membrane and membrane trafficking. However, their exact composition and function and the principles underlying their formation are only poorly understood. We employ biochemical, biophysical and cell biological methods to assess basic principles of membrane domain formation, visualize membrane domains in live mammalian cells and analyze their function in exocytotic and endocytic membrane transport.
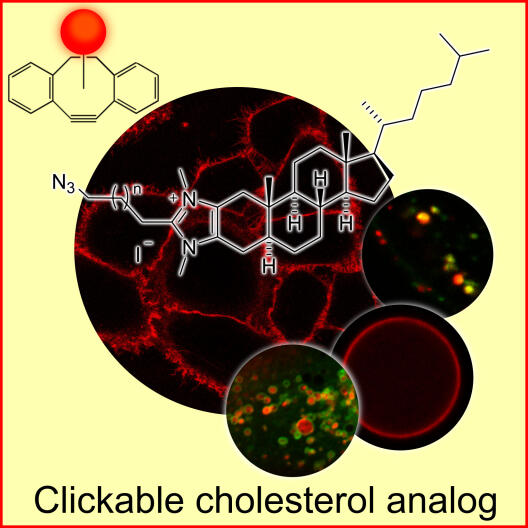
Among other things we collaborate with the groups of Bart Jan Ravoo and Armido Studer on the formation and characterization of novel redox-sensitive, cyclodextrin-based nanocontainers that we use to load fluorescently labeled lipids into cells. Lipid and sterol analogs that can be specifically labeled on the surface of cells and are synthesized in the group of Frank Glorius are also used to visualize the spatiotemporal distribution of these compounds in cellular membranes. Furthermore, properties of membrane lipids and membrane-binding proteins are studied in model membrane systems employing either vesicular structures such as giant unilamellar vesicles (GUV) or solid supported lipid bilayers (SLB).

