

Research Interests of the Molecular Evolution and Sociobiology Group
A major challenge for the future is to map genetic, genomic and epigenetic variation to observable phenotypic variation and understand how these traits evolve. We are using social (ants and bees) and solitary insects (parasitoid wasps like Nasonia) to understand the genetic basis and regulation of adaptive traits across multiple levels from gene to colony and time frames from seconds to millions of years.
1. Sociogenomics
Sociogenomics has two basic meanings, first it can refer to the study of genomes of social organisms or second it can be the study of the genomic basis of social behavior. We are interested in both and use genomic, transcriptomic, proteomic, and epigenetic methods to study the basis and structure of social genomes and social behavior.
Evolution of social traits in the ant genus Pogonomyrmex and Myrmecocystus. Pogonomyrmex and Myrmecocystus are New World genera that inhabit mesic and dry habitats in North and South America and North America, respectively. Currently about 100 species have been described for Pogonomyrmex and about 30 species for Myrmecocystus.

© Elizabeth Cash Fig.1: Pogonomyrmex barbatus worker at work (excavating and inspecting a non-nestmate)!

© Ti Eriksson Fig.2: Two Myrmecocystus mendax workers exchanging liquid food and part of a nest with queen and the famous honeypots (workers that have been converted to food storage containers).
In collaboration with RE Robertson (ASU) and Corrie Moreaux (Field Museum Chicago) we are generating a molecular phylogeny for Pogonomyrmex and working on improving the Myrmecocystus phylogeny we generated about 10 years ago. We have determined the social structure of several species in terms of mating frequency and queen number to understand the evolution and genetic and environmental basis of the observed variation in social structure in these ant genera. This also involves the study of two social parasitic species that have lost their worker caste and two species that evolved a derived genetic caste determination system. To elucidate the evolution of these two interesting variations of social behavior we use phylogeographic approaches.
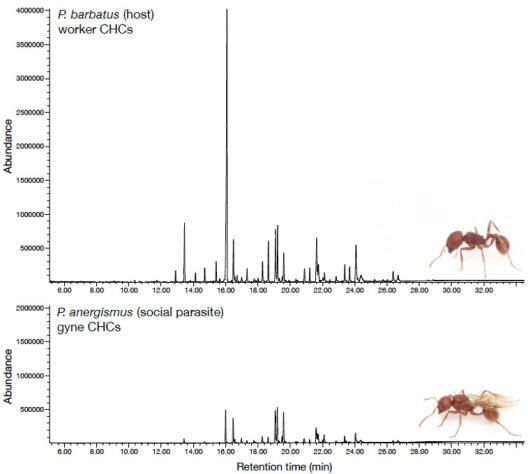
© AG Gadau Fig. 3: Comparison of the cuticular hydrocarbon profile of the social parasite P. anergism and a worker of its host P. barbatus. There are quantitative a series of qualitative differences.
One specific question we are currently trying to answer is the evolution and maintenance of primary polygyny that has evolved independently in Myrmecocystus mendax and Pogonomyrmex californicus. Most populations of these two species found their colonies as single queens and each mature colony has only one queen. However, both species have isolated populations that show primary polygyny, i.e. unrelated queens found a colony together and remain multiqueen societies. One of our studies is currently tackling the evolution and developmental regulation of aggression and cooperation/tolerance in Pogonomyrmex californicus.

© Elizabeth Cash Fig.4: Two Pogonomyrmex californicus queens from a primary polygynous population hanging out with larvae and pupae on a bed of seeds.
2. Evolution and genetic architecture of chemical communication
We use a combination of detailed phenotypic analysis (GC-MS, quantitative genetics (QTL analysis,qPCR), genetic manipulations (CRISPR/CAS, dsRNAi) to determine and confirm which genes/gene families are involved in chemical communication involved in colony, caste and species recognition. For example, we study the evolution of gene families involved in chemical communication (desaturase, olfactory and chemical receptor proteins, etc.) in insects in general and Pogonomyrmex and members of a parasitoid wasp genus Nasonia spp. in particular.
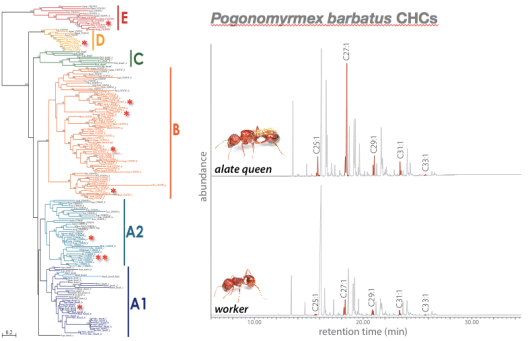
© AG Gadau Fig. 5: Evolution of Desaturase genes (left) and cuticular hydrocarbon profile differences between workers and alate queens of P. barbatus (alkene peaks are highlighted). Alate queens have a significantly higher concentration of alkenes in their profile than workers from the same colony. It is not known which of the 11 desaturase genes in P. barbatus are responsible for this difference.

Nasonia viritipennis female on micro array© AG Gadau Nasonia viritipennis
Nasonia is the “other” Hymenopteran model organism besides the honeybee. Nasonia spp. has a generation time of app. 14 days and we can produce hybrids in the laboratory once the lines are cured from its endoparasite Wolbachia, which otherwise causes cytoplasmic incompatibility. In Nasonia we can map, identify and confirm almost any interspecific variable trait within 6-12 month. So far we mapped male wing and body size differences, male courtship behavior, components of the cuticular hydrocarbon profile, developmental traits (craniofacial anomalies), nuclear-cytoplasmic incompatibilities, etc..
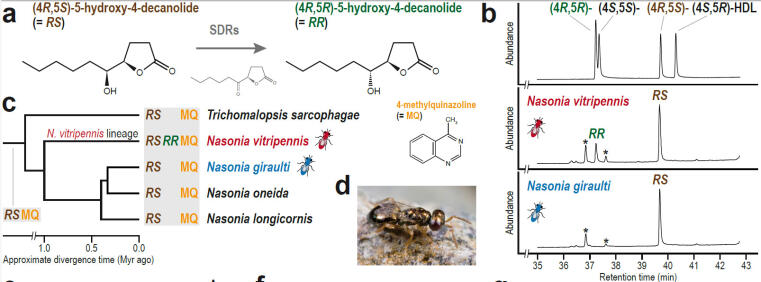
© AG Gadau Fig. 6: Evolution of a novel male sex pheromone in Nasonia vitripennis. The male sex pheromone of N. vitripennis has an additional component RR that females of use to distinguish between own and allospecific males of the sympatric N. giraulti. (For details see Niehuis et al.2013)
Nasonia males differ significantly in their cuticular hydrocarbon profile (also used as species and sex signal) and we are using hybrid crosses to map QTL and eventually identify the genes underlying those differences.

© AG Gadau Fig. 7: GC-MS profile of males from N. vitripennis and N. giraulti (left) and overlap of QTL for differences in individual cuticular hydrocarbon peaks and candidate genes (desaturases and elongases) based on hybrid males (right).
3. Speciation – Mechanisms and Processes
Pre- and Postzygotic isolation - Under the biological species concept, speciation is the evolution of hybridization barriers. We have a long-term interest in discovering genes underlying pre- and postzygotic isolation in Nasonia spp. (e.g. pheromones (s. Fig. 6 and 7), nuclear-cytoplasmic incompatibilities, microbiome versus genome) and their evolutionary history.
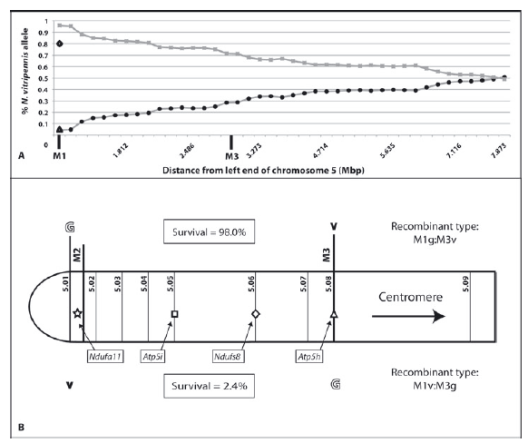
© AG Gadau Fig.8: (A) Percentage of Nasonia vitripennis alleles along the distal part of the left arm of chromosome 5 in N. vitripennis ×N. giraulti F2 hybrid males with N. giraulti cytoplasm ( = VG[G]). (B) Distal part of left arm of chromosome 5. The survival rate of the specific type of recombinant individuals is given in enclosed boxes as observed percentage of total expected number of individuals of each recombinant type. Symbols in the chromosome represent candidate genes described here (star = Ndufa11) and from Gibson et al. (2010) (square, Atp5i; diamond = dufs8; and triangle, Atp5h).

