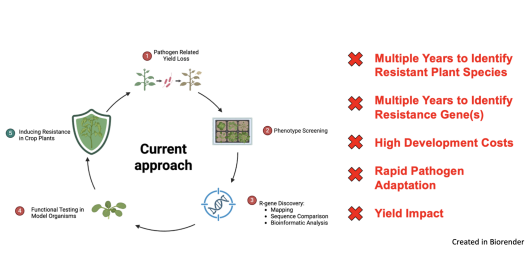
Our mission is to break this vicious cycle. We strive for modular, broad-spectrum resistance, not once it’s too late, but ahead of time. To achieve this, we focus on conserved virulence factors of pathogens. Many crop pathogens across kingdoms express conserved proteases as key virulence factors in plant cells.
Hereby, conserved proteases are deployed on two levels:
Agriculturally relevant RNA viruses (such as poty-, seco-, and closteroviruses) express virus-encoded proteases that process polyproteins to properly assemble new viruses.
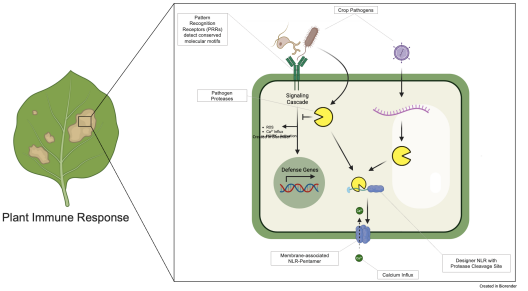
Bacteria and fungi are detected by Pattern Recognition Receptors (PRRs) via conserved molecular motifs, such as flagellin and chitin. To mask their presence, proteases are deployed that disrupt signaling cascades at key levers. This grants the pathogen its most valuable resource: Time to multiply. As such, the most effective treatment is an early warning system.
Plant nucleotide-binding leucine-rich repeat receptors (NLRs) regulate immunity and cell death. Often occurring in pairs, many primary NLRs recognize a specific effector and activate a secondary RPW8 domain-containing “helper” NLR, such as the Arabidopsis NRG1.1 protein. When activated, secondary NLRs oligomerize and form a selective calcium ion channel, irreversibly inducing an immune reaction, visible as a hypersensitive reaction (HR).
Prior studies have demonstrated that a single nucleotide substitution in the AtNRG1.1 gene can generate an autoactive NLR form. As the functionality relies on negatively charged N-terminal residues, the autoactivity can be masked by fusing an uncharged peptide (blocking linker) to the N-terminus.
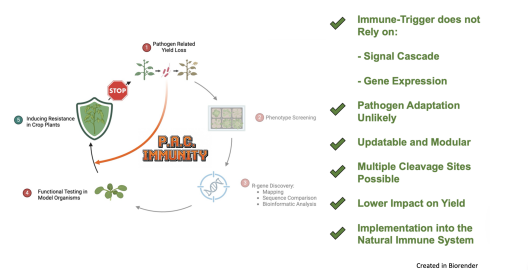
When the N-terminal blocking linker contains a protease-specific cleavage site, this minimal setup becomes a designer immune receptor. Specific cleavage by a pathogen protease reveals the functional N-terminus, subsequently inducing an immune response. As the only variance in this setup is the cleavage site, it becomes a “print-on-demand system” that can be updated, if necessary. Not by searching for an entirely new gene, but by simply switching the site of interest (Fig. 2).
The functionality and modularity of this framework were successfully demonstrated, conferring broad-spectrum resistance against multiple potyviruses in the model organism (Nicotiana tabacum), as well as in soybean field tests.