Epigenetics meets metabolism
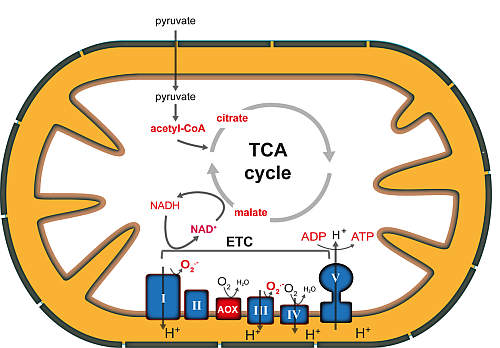
The energy-demand and metabolic requirements of plant cells differ under changing environmental conditions. Mitochondria and chloroplasts, the cellular organelles of respiration and photosynthesis, respectively, are central hubs in the conversion of energy and redox homeostasis in plant cells. They are connected to metabolic pathways from different subcellular compartments. Both organelles are ideally placed to act as sensors of the energetic and metabolic status of the plant cell. Perturbations of the cellular energy status can lead to a reconfiguration of organellar activities, which in turn have profound effects on other cellular compartments, including nuclear gene expression. Changes in nuclear gene expression triggered by signals derived from metabolic perturbations in organelles are termed retrograde responses and are critical to allow adaptation of the bioenergetic state of the organelles. Nearly nothing is known about the signaling processes transducing the retrograde responses. We are interested in deciphering the role of post-translational modifications (PTMs) pf proteins in this process. PTMs provide a powerful mechanism to rapidly and temporarily alter protein functions and locations in the cell, and they are also capable of providing information to regulatory proteins by creating docking sites for PTM-recognition domains. Intriguingly, the substrates of several major PTM-modifying enzymes are either central metabolites or redox-active compounds, as for example ATP, acetyl-CoA, NAD+, and glutathione, which suggests that the activity of these enzymes could be directly coupled to the energy status or redox status of the of the organelle. The main goal of our research is to unravel the regulatory processes that coordinate organellar functions in dependence on environmental and metabolic cues in different plant tissue.
Regulation of protein functions by acetylation
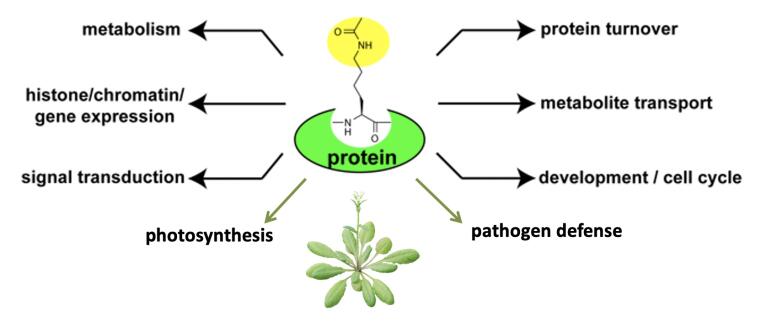
Reversible acetylation of the ε-amino group of lysine has recently emerged as a major post-translational modification of proteins controlling many important cellular functions beyond histone modification. Acetylation of lysine residues of proteins is regulated by the activities of acetyltransferases and deacetylases. Both our own recent results in Arabidopsis and research in animal systems showed that lysine acetylation acts as an on/off switch for enzyme activities as well as a signal for cargo transport in several cellular processes regulating energy metabolism, signaling cascades and cytoskeleton dynamics. The aim of our research is to reveal the function and importance of lysine acetylation in the regulating of plant metabolic pathways and signaling processes involved in the plant stress response and its cross-talk with other PTMs. We are investigating thefunctions of the diverse family of histone / lysine deacetylases (HDAC/ KDAC) and histone / lysine acetyltransferases (HAT / KAT) in this process.
Sensing and signaling of metabolic states in plants
One of the most important mechanisms to coordinate adjustments of the metabolic network is by regulation via internal signals generated from specific metabolites to regulate gene expression in the nucleus. A large number of metabolic genes is already known whose expression is regulated by altered concentrations of key nutrient metabolites such as sugars and nitrate. Furthermore, evidence is emerging that metabolite signaling of gene expression may be mediated through many additional metabolites, such as organic acids. The aim of our research is to identify the role of central energy metabolites in the regulation of cell functions.